Dual anti-platelet therapy (DAPT), consisting of a P2Y12 inhibitor and aspirin, is the standard of care following acute coronary syndrome (ACS) with or without percutaneous coronary intervention (PCI), and can reduce major adverse cardiovascular events (MACE) and post-procedural atherothrombotic events when used for 12 months.1,2
Ticagrelor and prasugrel are recommended over clopidogrel in international guidelines owing to their greater potency and consistency in platelet inhibition.1,3,4 Clopidogrel is subject to inter-individual variability resulting from the perturbations of a two-step bioactivation, thought to be attributed to the genetic polymorphisms of the cytochrome P450 2C19 (CYP2C19) gene.5 Individuals with CYP2C19*2 and/or CYP2C19*3 loss-of-function (LoF) alleles are expected to have reduced clopidogrel bioactivation, leading to inadequate platelet inhibition or high platelet reactivity (HPR), which is associated with an elevated risk of MACE.6–8 CYP2C19 polymorphism is more prevalent in Asian than in white populations. The prevalence of CYP2C19 LoF in Singapore has been reported to be as high as 62%.9 Nevertheless, clopidogrel remains clinically relevant for its wider generalisability of use, affordability and lower risk of bleeding.10
Beyond CYP2C19 polymorphism, clinical risk factors have been shown to significantly contribute to clopidogrel hypo-responsiveness.11 The novel ABCD-GENE score was devised as a screening tool to identify potential patients with on-clopidogrel HPR, who were at higher risk of MACE. Angiolillo et al. found that a 10-point cut-off predicted 1-year risk of all-cause mortality, a composite of all-cause death, MI and stroke and moderately discriminated HPR in a largely American cohort.12 The 10-point cut-off was later re-established by Capodanno et al., whereas Saito et al. instead suggested that a 9-point cut-off best predicts on-clopidogrel HPR in Japanese patients.13,14
To date, most studies have used the VerifyNow P2Y12 test (VerifyNow) to assay clopidogrel-induced platelet reactivity.12,14 The influence of other point-of-care assays, such as multiple electrode aggregometry (MEA), on the ABCD-GENE score is not as well studied.15 The aim of this study was therefore to validate the prognostic utility of the ABCD-GENE score by characterising a best cut-off to, first, predict the 1-year incidence of MACE and second, predict on-clopidogrel HPR in our cohort of Asian ACS patients.
Methods
Study Design
The study population was derived from the P2Y12 inhibitor in Acute Coronary Syndrome (PACS) registry, a prospective study that enrolled ACS patients to primarily evaluate the relationship between MEA cut-offs with MACE and bleeding.
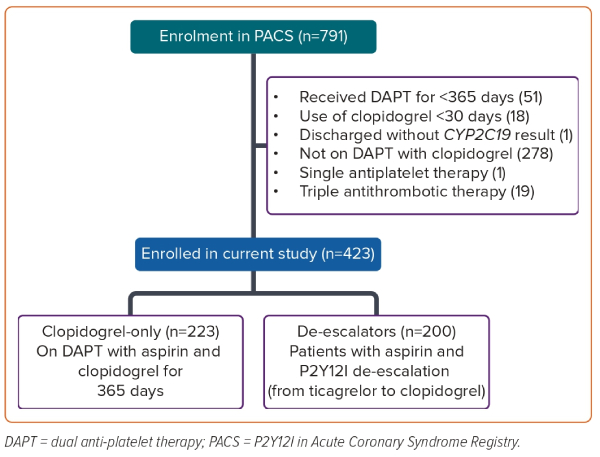
Patients in this post hoc analysis received DAPT for ACS between 2011 and 2019. Use of DAPT was defined as the combined prescription of aspirin and either clopidogrel or ticagrelor. Patients who used either single antiplatelet or triple anti-thrombotic therapy (defined as the concurrent use of oral anticoagulant with DAPT), received DAPT for less than 365 days or did not have a valid CYP2C19 genotype were excluded (Figure 1).
Patients who completed 365 days of DAPT but underwent P2Y12 inhibitor de-escalation were classified as ‘de-escalators’. For the patients who underwent de-escalation, the duration of DAPT on clopidogrel was calculated using the difference (in days) between the intended DAPT end date and the de-escalation date. Patients were excluded if the switch occurred in the final month of the intended DAPT duration, or if there was more than one switch in P2Y12 inhibitor. Patients who underwent P2Y12 inhibitor escalation were excluded because they had all received <30 days of clopidogrel. Patients who completed all 365 days of DAPT on clopidogrel were classified as the ‘clopidogrel-only’ group. All eligible patients were routinely followed from the point of discharge until the first outcome of interest or up to 12 months from the date of P2Y12 inhibitor initiation, whichever was earlier.
Given that all of the patients enrolled in PACS consented to further research, additional approval from the institutional review board was not required. PACS was conducted in accordance with Good Clinical Practice Guidelines and the principles of the Declaration of Helsinki.
ABCD-GENE Score Calculation
The ABCD-GENE score for each patient was tabulated by allocating corresponding points when any of the components were present.12 Four points were allocated for age >75 years or BMI >30 kg/m2, 3 points were allocated for stage 3–5 chronic kidney disease or diabetes, and 6 and 24 points were allocated if one or two CYP2C19 LoF alleles were present, respectively. For the analysis of the correlation with MACE, patients were stratified into two ABCD-GENE score groups: below 10 points (the <10 group) or 10 points and above (the ≥10 group).
Clinical Outcomes
The primary objective was to determine the cut-off that best predicted MACE at 1 year. A score range of 7–10 was used to determine the best cut-off for MACE in our study.14 MACE was defined as a composite event of all-cause mortality, MI and repeat revascularisation.16 All ischaemic events were independently adjudicated by a cardiologist blinded to the treatment assignment in the PACS study.
The secondary objective was to identify the best cut-off that predicts on-clopidogrel HPR measured using MEA. Patients with MEA ≥46 U were classified as HPR, and those with MEA <46 U were classified as non-HPR.17 Patients were also stratified into the respective subgroups, either the clopidogrel-only or the de-escalator groups, to ascertain whether the duration of clopidogrel use influenced HPR prediction by ABCD-GENE score. Sensitivity analyses were carried out to ascertain the robustness of the ABCD-GENE score (Supplementary Table 1).14 Variables for sensitivity analysis were chosen using the baseline characteristics that were significantly different between the HPR and non-HPR groups (Supplementary Table 2), and according to the literature and expert opinion.
Statistical Analysis
Continuous variables were assessed for normality using the Kolmogorov–Smirnov test. Differences were assessed using the independent samples t-test for parametric variables or the Mann–Whitney U-test for non-parametric variables, and reported as either mean ± SD or median (IQR), respectively. Ordinal variables were analysed using the Mann–Whitney U-test, and nominal variables were analysed either with the chi-squared test or Fisher’s exact test. Both ordinal and nominal variables are described as frequency and proportion (n (%)).
The relationship between the ABCD-GENE score and MACE was ascertained with model 1 and the adjusted model 2 using Cox regression, via backward stepwise elimination, with an entry and exit p-value of 0.10 and 0.05, respectively. Variables used for the adjusted model were selected using univariable Cox regression (Supplementary Table 3), relevant publications and expert opinion. The ability of the ABCD-GENE score to predict on-clopidogrel HPR was assessed using the area under the receiver operating characteristic (ROC) curve (AUC). The sensitivity and specificity were generated via cross-tabulation analysis and the best cut-off was identified using Youden’s index.18
A priori sample size calculation was not performed for this explorative study. All statistical analyses were performed as a two-tailed test with 95% CIs, using IBM SPSS Statistics 27.0.
Results
Baseline Characteristics
A total of 423 subjects were eligible for the validation study, with 223 and 200 in the clopidogrel-only and de-escalator groups, respectively. While all subjects in the clopidogrel-only group received clopidogrel for 365 days, the de-escalator group received clopidogrel for a median of 236 days (range: 167–270 days; Supplementary Figure 1).
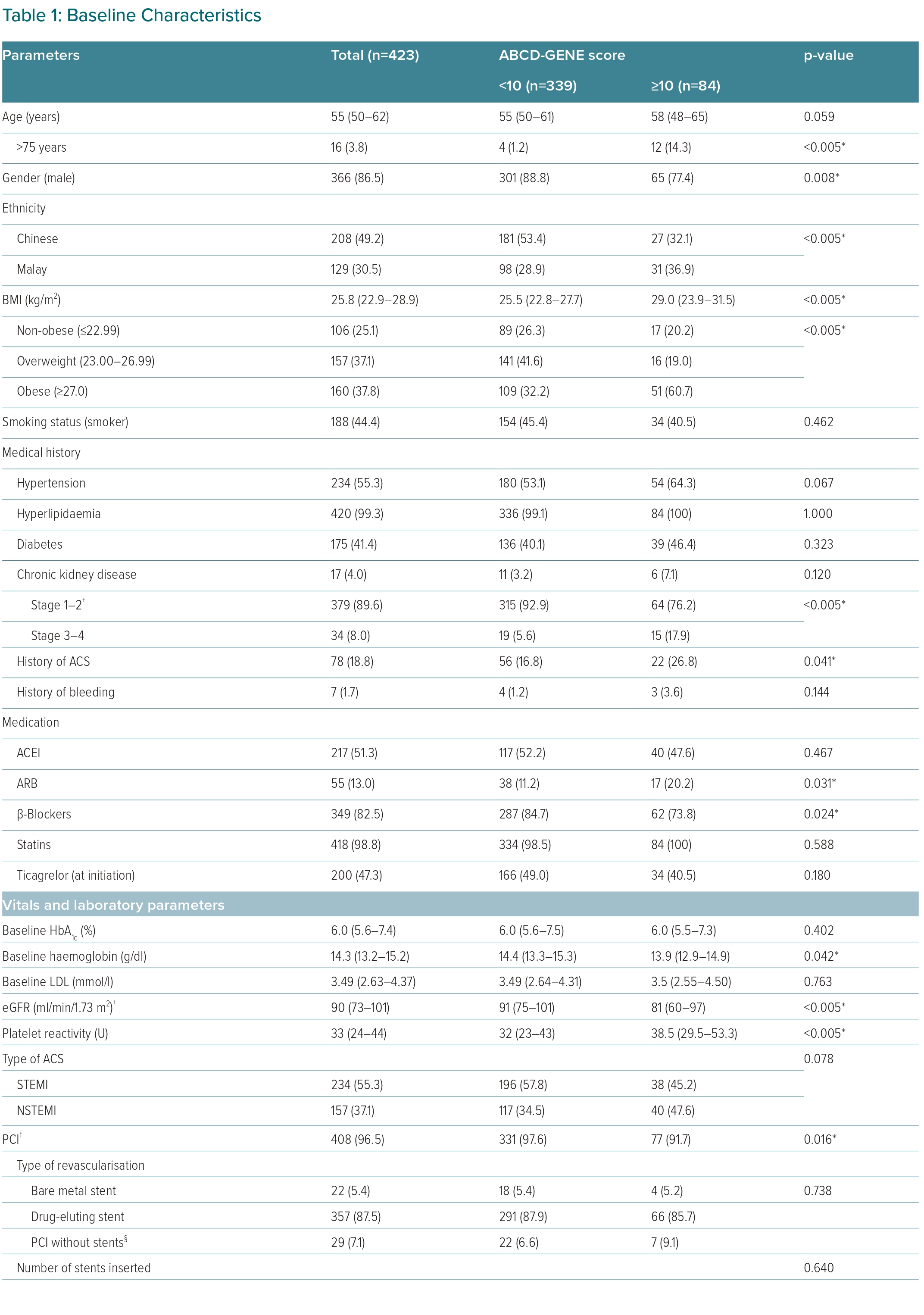
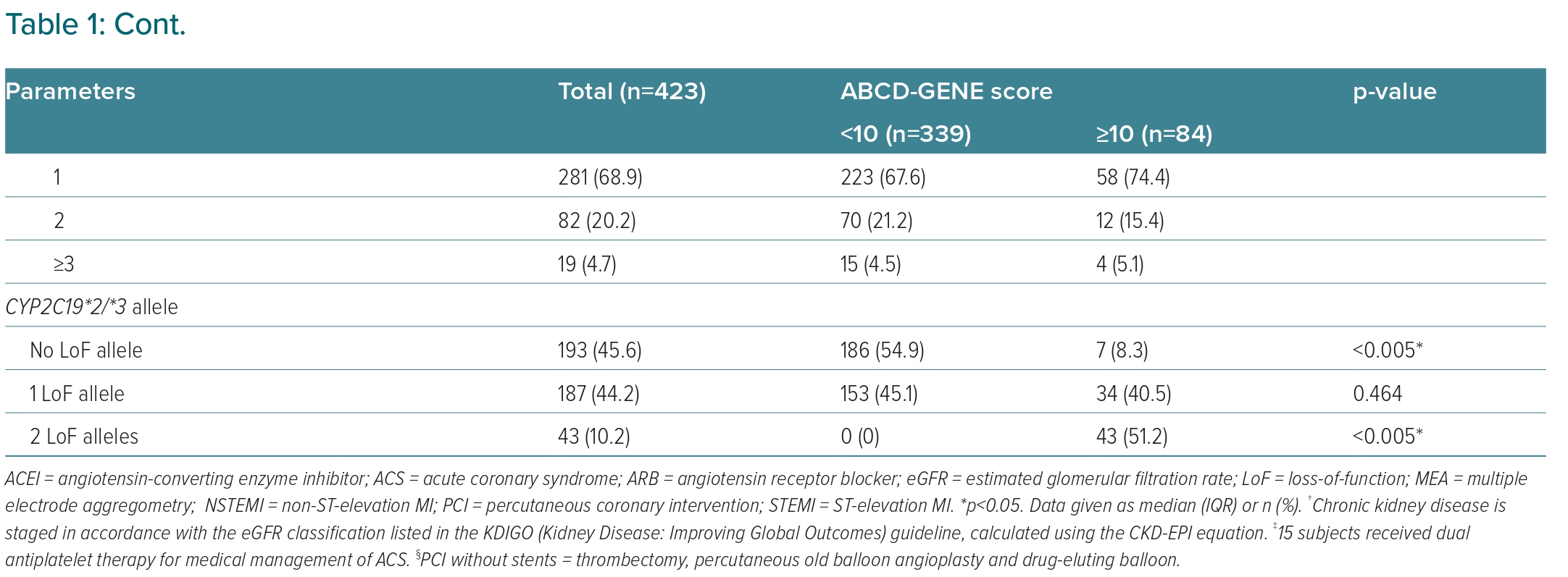
Altogether, 84 patients had ABCD-GENE score ≥10 points and 339 patients had ABCD-GENE score <10 points (Table 1). Notably, ethnic group, female sex, history of ACS, higher baseline LDL and haemoglobin, higher platelet reactivity when treated with clopidogrel, not receiving PCI and hospital discharge with angiotensin receptor blocker (ARB) or β-blocker use were associated with the higher ABCD-GENE score of ≥10 points in our cohort.
Best Cut-Off for Predicting the Incidence of MACE at 1 year
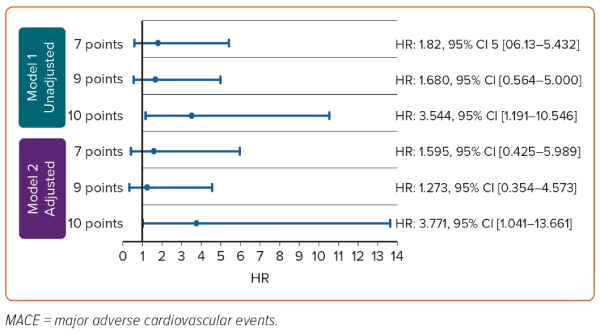
At 1 year, six patients (7.1%) with ABCD-GENE score ≥10 points and seven patients (2.1%) with ABCD-GENE score <10 points had MACE, respectively. The dichotomous classification using 10 points as the cut-off independently predicted the 1-year risk of MACE (Supplementary Table 4), and with a high accuracy of 80%. Patients with a score of ≥10 points were almost fourfold more likely to develop MACE than those with a score of <10 points regardless of adjustment (unadjusted HR 3.544; 95% CI [1.191–10.546]; adjusted HR 3.771; 95% CI [1.041–13.661]). Sex, history of ACS, baseline LDL and ARB use were also independently associated with the 1-year risk of MACE. A subgroup analysis of only PCI patients noted a similar outcome (HR 3.696; 95% CI [1.007–13.569]) using the 10 point cut-off (Supplementary Table 5).
A sensitivity analysis was carried out to test all other cut-off scores, and only the cut-off at 10 points was able to differentiate the 1-year risk of MACE. The relationship between the candidate cut-offs and MACE is shown in Figure 2.
Best Cut-Off to Predict On-clopidogrel HPR
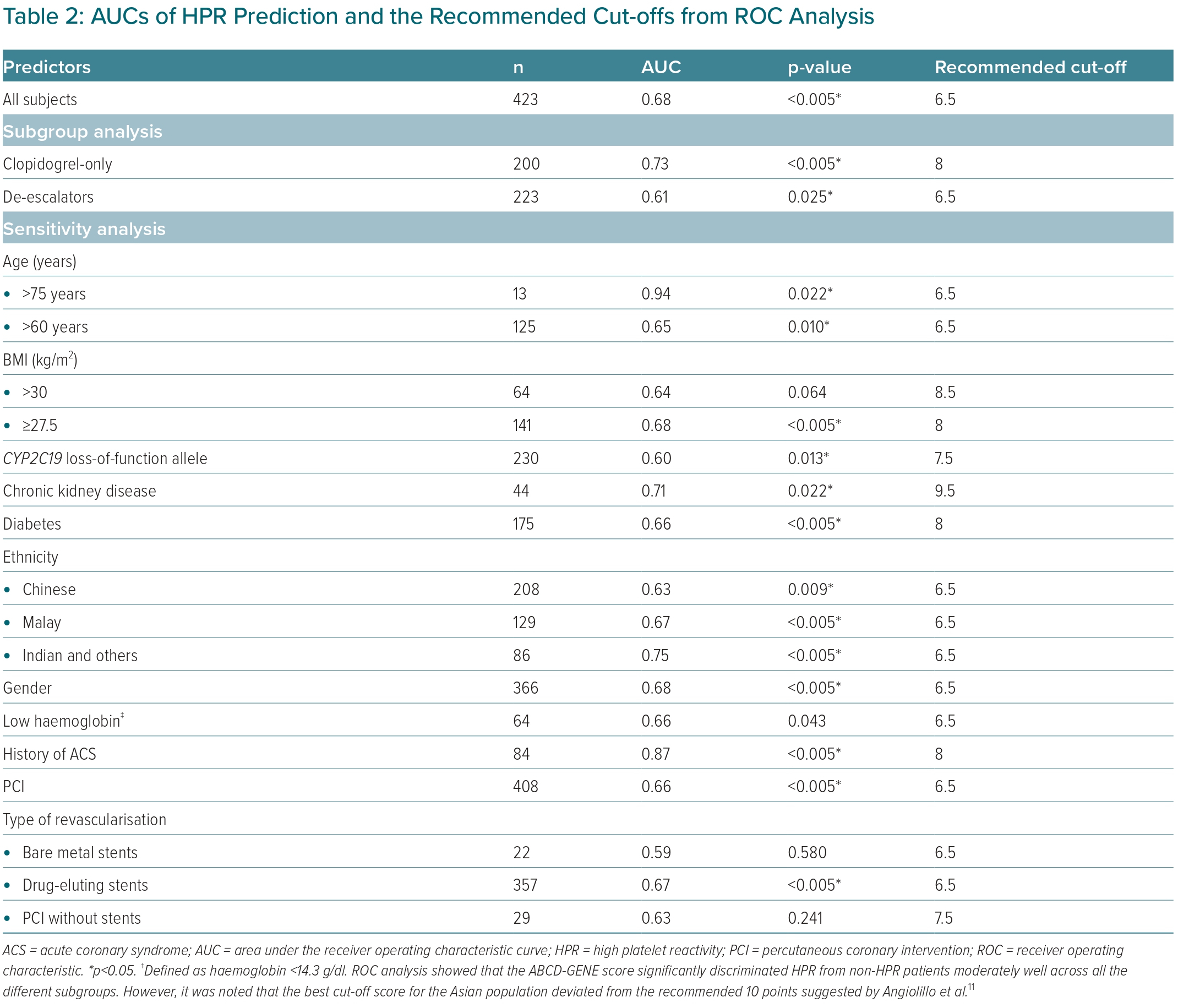
Using ROC analysis, the ABCD-GENE score had moderate prognostic accuracy to predict on-clopidogrel HPR (AUC 0.68; p<0.005; Table 2). A greater prognostic accuracy was observed in the clopidogrel-only subgroup (AUC 0.73; p<0.005) than the de-escalator subgroup (AUC 0.61; p=0.025) (Supplementary Figure 2). Additionally, the score was able to discriminate between HPR and non-HPR across sensitivity analysis, except for those with BMI >30 kg/m2 (AUC 0.64; p=0.064) and those who received bare metal stents (AUC 0.58; p=0.580) or PCI without stents (AUC 0.63; p=0.241).
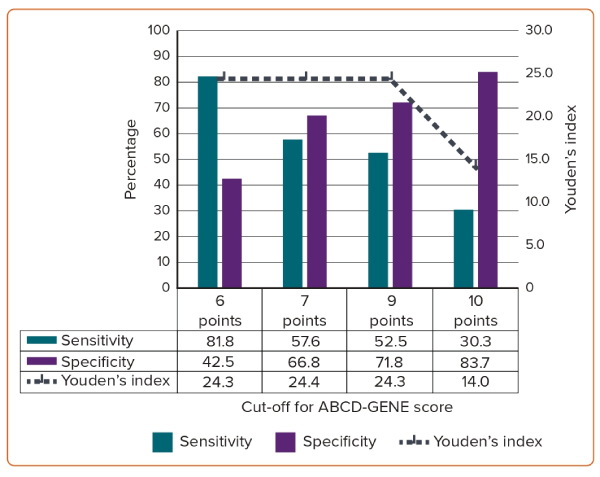
Across all analyses, the recommended cut-off scores consistently ranged between 6.5 points and 9.5 points, and repeatedly demonstrated specificity but poor sensitivity in predicting HPR. As the score increases from 6 points to 10 points, the sensitivity decreases from 81.8% to 30.3%, while specificity increases from 42.5% to 83.7% (Figure 3). Using Youden’s index, 7 points was the ideal ABCD-GENE score cut-off to predict on-clopidogrel HPR.
Discussion
Our study retrospectively validated the prognostic utility of the ABCD-GENE score by identifying the best cut-off that predicted MACE and on-clopidogrel HPR. The key findings from our study were that a 10 point cut-off moderately predicted the 1-year risk of MACE, and that the adjusted 7 point cut-off satisfactorily predicts HPR but does not predict MACE well.
Patients with ABCD-GENE score ≥10 points were fourfold more likely to have MACE than those with a score <10 points, comparable to the 1.5-fold and twofold increased composite risk of all-cause mortality, MI and stroke in largely white cohorts.12,13 The greater risk of MACE in Asian patients than in white patients was hardly surprising, given that the prevalence of CYP2C19 LoF is almost twice that in Asian than in white populations.9
In our adjusted model, we also identified additional factors associated with MACE, including baseline LDL, female sex, and history of ACS and ARB use. Notably, every 0.1 mmol/l increase in baseline LDL elevates MACE risk by 89% in our cohort. This trend was consistent with a previously reported association in which uncontrolled LDL levels consequentially exacerbate cardiovascular events.19,20 However, all of the present patients had been put on potent statins following the ACS, and this serves to remind clinicians of the residual risk of neglecting LDL control even after the commencement of statins. Asian female patients were previously found to be poor responders to clopidogrel, and those with a history of ACS were likely to encounter a higher baseline platelet reactivity.21 Both factors contributed to clopidogrel hypo-responsiveness, increasing the likelihood of MACE.11 Interestingly, the use of ARBs was the strongest predictor of MACE. Although ARB use has been associated with a higher rate of MACE and revascularisation than angiotensin-converting enzyme inhibitor (ACEI) use, it is plausible that this observation was a result of chance; the validity of ARB use in the association with MACE should be ascertained in subsequent studies.22,23
The ABCD-GENE score also significantly and moderately distinguished HPR from non-HPR. At an AUC of 0.68, the overall discernment was comparable with the original validation cohort (AUC = 0.64), but was weaker than that for the Japanese cohort (AUC = 0.78).12,14 A moderate-to-strong discrimination was observed across the sensitivity analysis, except for only a few factors: BMI >30 kg/m2, use of bare metal stents and PCI without stents. Considering the lower BMI cut-off for obesity of 27.5 kg/m2 in Asia, the use of a BMI cut-off of >30 kg/m2 may not be ideal.24 Similarly, a reduced group of patients receiving bare metal stents or PCI without stents may limit the internal validity of the association between the score and HPR. Both may contribute to the non-significant HPR predictions.
The best cut-off at 7 points is lower than the 9 points identified in the Japanese cohort.14 However, the HPR predictive utility in the present cohort consistently ranged between 6.5 points and 9.5 points across ROC analyses, similar to the Japanese cohort, while the cut-off using 10 points performed poorest across the candidate cut-offs, although the high specificity and low sensitivity trend was similar to that observed in the white cohort.12,14
The discordance in cut-off points for MACE and HPR prognostication suggests a lack of association between MEA-determined HPR and actual MACE. As opposed to VerifyNow in published validations, the platelet reactivity in the present cohort was quantified using MEA.12,14 Although both assays are favoured for their efficiency in measuring whole blood, the results lack agreement and are unequally effective in predicting clinical outcome.15,25 For ACS–clopidogrel–HPR patients, a significantly stronger relationship has been observed between VerifyNow and MACE than for MEA.26,27 This was also observed in the present analysis, in which the inclusion of neither MEA measurements nor platelet reactivity status influenced the effect estimates of MACE prediction at 10 points. Although MEA might be less sensitive in predicting adverse ischaemic events than VerifyNow, the use of unvalidated cut-offs for HPR in a predominantly Asian population, may not accurately reflect the actual haemostasis, resulting in the misclassification of HPR status against MACE.17 Furthermore, the present underpowered study (with only 13 subjects diagnosed with MACE) also limits the effectiveness of MEA.
The value of DAPT use is to reduce recurrent MACE while HPR is merely a surrogate. Clinicians will find it more intuitive to use clinical factors coupled with CYP2C19 phenotypes. This would also avoid inter-population variations in platelet reactivity cut-offs, and clinical variability arising from the use of platelet function analysers.25,26 Altogether, greater credence should be given to the ability of the ABCD-GENE score to predict MACE than to identify HPR. As such, the evidence thus far supports the use of the ABCD-GENE cut-off at 10 points in our Asian cohort.
Compounded by the pharmacological and clinical complexities surrounding DAPT, the ischaemic–bleeding conundrum, characterised by the concurrent need to achieve anti-ischaemic activity without elevating bleeding risk, suggests that the use of ticagrelor over clopidogrel may not always be clinically feasible.28,29 Instead, the need to individualise P2Y12 inhibitors for ACS patients raises interest in personalised DAPT. Currently, the applicability of platelet reactivity-guided DAPT remains apocryphal, and head-to-head trials on genotype-guided DAPT have produced mixed outcomes.7 The observed prognostic utility of the ABCD-GENE score may be useful to improve the effectiveness of individualised DAPT for ACS patients.
To the best of our knowledge, our study is the first in Asia to validate the ABCD-GENE score cut-off for both MACE prognostication and on-clopidogrel HPR with real-world evidence since the study by Angiolillo et al.12 The identified best cut-off in our heterogeneous cohort is generalisable to southeast and central Asian countries, for instance, China, Malaysia and Indonesia. In agreement with recent findings, this also suggests that CYP2C19 genotyping would be relevant as a guide for DAPT in the region.30
However, our study has some limitations. First, the use of MEA in our study limits comparison between our cohort and published studies, given that most studies used VerifyNow.12,14 However, MEA was the only assay available to the PACS investigators because VerifyNow was not registered for use in Singapore. Furthermore, there is no gold standard established for HPR.15 Nonetheless, we believe that the ability of the ABCD-GENE score to predict MACE is more important than for HPR.
Additionally, our findings should be interpreted with caution. Relative to the 14.3% incidence of MACE in the original validation, the smaller incidence of MACE in the present cohort (3.1%) potentially widens the confidence interval for MACE prediction, reflecting a likelihood of false negatives (i.e. type II error), and consequently leading to a low 46.2% sensitivity.12 The impact of ticagrelor use in the de-escalator group on 1-year risk of MACE was not studied due to the small sample size, which limited the clinical event count.
Conclusion
The ABCD-GENE score was able to predict MACE with a high accuracy in our cohort, indicating the potential value of the risk model. Given that the present study was hypothesis generating in nature, further studies in an expanded cohort of Asian ACS patients using the ABCD-GENE score in addition to the newly described predictors, are warranted.
Click here to view Supplementary Material
Clinical Perspective
- The ABCD-GENE score was devised as a screening tool to identify potential patients with on-clopidogrel high platelet reactivity (HPR), who were at higher risk of major adverse cardiovascular events (MACE).
- The original 10 point cut-off predicted the 1-year risk of all-cause mortality, a composite of all-cause death, MI and stroke and moderately discriminated HPR in a largely American cohort.
- This cut-off score was reduced to 7 points in an Asian population, with new predictors identified such as female sex, baseline LDL, history of acute coronary syndrome (ACS) and use of angiotensin receptor blockers.
- Further studies in an expanded cohort of Asian ACS patients using the ABCD-GENE score in addition to the newly described predictors are warranted to evaluate the clinical applicability and generalisability of this screening tool beyond the largely American population.